JustClust
What is JustClust?
JustClust is a tool for analysing biological data with cluster analysis. JustClust can handle many formats of data and cluster the data with many state-of-the-art techniques. The aim of JustClust is to provide an easy-to-use application which can perform any analysis on any data.
Overview
Most algorithms for clustering biological data have their own software just for that algorithm. If a biologist wants to use multiple algorithms, they will have to download the software for each algorithm and learn to use it. JustClust takes many algorithms which are used frequently in biologicaly and puts them all in one place for ease of use. JustClust is itself also very easy to use with intuative controls.
JustClust currently includes the affinity propagation, average-linkage, Chinese Whispers, ClusterONE, complete-linkage, connected components, DBSCAN, MCL, MCODE, single-linkage, and SPICi clustering algorithms. The file formats supported by JustClust are those of fasta files, microarray files, tab-separated values files, comma-separated values files, and GML files.
JustClust displays its results with graphs. The layout of these graphs can be altered by the user and there are also a number of built-in layout functions which can lay the graphs out automatically. These include the prefuse force layout and the Cytoscape circular, degree sorted circle, edge-weighted spring embedded, and grid layouts.
The clustering algorithms, file parsers, and graph layouts are all plug-ins and JustClust can be extended easily with more plug-ins of these types which have been developed by users.
Plug-ins
We welcome new plug-ins for the parsing, clustering, and visualisation of data in JustClust. The Plug-in Development Manual describes in detail how plug-ins can be developed.
If you have developed a plug-in, we will be glad to include it in the next release of JustClust. Please send it to support-at-paccanarolab.org.
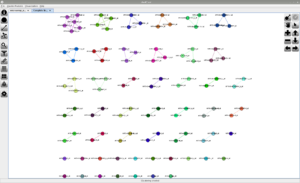
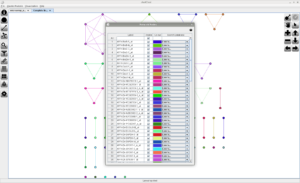
Screenshots
Click on any of the images below to get an idea of how JustClust looks and whether it is suitable for you.
Download
Zip file with installer + datasets used in the paper
Documentation
User Manual
Download
Plug-in Development Manual
Download
Get the source code
JustClust is open-source. If you are interested in developing it further, feel free to get the source code from GitHub and experiment with it. You will need Git to check out the source code.